Cleaning data with sqlite-utils and Datasette
This tutorial will show you how to use the sqlite-utils command line tool import data into SQLite, clean it and then explore it with Datasette.
The concepts in this tutorial are also demonstrated in this video of a talk I gave at the HYTRADBOI conference in April 2022.
Start with some CSV data
The Florida Fish and Wildlife Conservation Commission maintain a database of manatee carcass recovery locations, with details dating back to April 1974.
You can request the latest copy as a CSV file from their website - or download this copy of the CSV: Manatee_Carcass_Recovery_Locations_in_Florida.csv generated on July 31st 2022.
Install sqlite-utils
Here are the installation instructions. If you are on a Mac with Homebrew installed you can run brew install sqlite-utils. Python users can run pip install sqlite-utils.
Import the CSV into a SQLite database
sqlite-utils is a command line tool, so you'll need to open up a terminal on your computer and change directory to the place where you saved the CSV file.
You can create a new manatees.db SQLite database like this:
sqlite-utils insert manatees.db locations \
Manatee_Carcass_Recovery_Locations_in_Florida.csv --csv -d
This sqlite-utils insert command inserts data into a database table, creating the table if necessary.
manatees.dbis the SQLite database file we are using - this will be created if it doesn't exist yet.locationsis the name of the table we are creatingManatee_Carcass_Recovery_Locations_in_Florida.csvis the CSV file we are importing--csvtells the tool to treat the incoming data as CSV data- The
-doption tells the tool to automatically detect the types of data. Without this, every column will be treated as text. With this option columns that have numeric data in will be used to populate numeric columns.
Having created the database, we can use the schema command to view its schema:
sqlite-utils schema manatees.db
The output of that command looks like this:
CREATE TABLE "locations" (
[X] FLOAT,
[Y] FLOAT,
[OBJECTID] INTEGER,
[FIELDID] TEXT,
[REPDATE] TEXT,
[REPYEAR] INTEGER,
[REPMONTH] INTEGER,
[REPDAY] INTEGER,
[SEX] TEXT,
[TLENGTH] FLOAT,
[STATE] TEXT,
[COUNTY] TEXT,
[LAT] FLOAT,
[LONG_] FLOAT,
[DCODE] INTEGER,
[MORTALITY] TEXT,
[created_user] TEXT,
[created_date] TEXT,
[last_edited_user] TEXT,
[last_edited_date] TEXT
);
The locations table here has been created with columns that match the columns that were present in the CSV file. The columns have the appropriate types thanks to the -d option we passed to sqlite-utils insert.
We can use the sqlite-utils tables command to see those tables:
sqlite-utils tables manatees.db --counts
Output is:
[{"table": "locations", "count": 13568}]
The --counts option adds a count of the rows in each table. That's over 13,000 deceased manatees!
By default, the tables command outputs JSON. You can change this to be a table format using the --table or -t options:
sqlite-utils tables manatees.db --counts -t
table count
--------- -------
locations 13568
Querying the data
Let's run a SQL query against your data. The sqlite-utils command to do that looks like this:
sqlite-utils manatees.db "select * from locations limit 2"
This outputs JSON, showing the first two rows and every column (thanks to that select *):
[{"X": -80.1176098026034, "Y": 26.1004059191504, "OBJECTID": 149433, "FIELDID": "M7401", "REPDATE": "1974/04/03 00:00:00+00", "REPYEAR": 1974, "REPMONTH": 4, "REPDAY": 3, "SEX": "F", "TLENGTH": 260.0, "STATE": "FL", "COUNTY": "Broward", "LAT": 26.100401, "LONG_": -80.117607, "DCODE": 9, "MORTALITY": "Undetermined: Other", "created_user": "FWC", "created_date": "2021/10/15 11:30:55+00", "last_edited_user": "FWC", "last_edited_date": "2021/10/15 11:30:55+00"},
{"X": -80.1664354041335, "Y": 25.8337161801256, "OBJECTID": 149434, "FIELDID": "M7402", "REPDATE": "1974/06/27 00:00:00+00", "REPYEAR": 1974, "REPMONTH": 6, "REPDAY": 27, "SEX": "M", "TLENGTH": 290.0, "STATE": "FL", "COUNTY": "Miami-Dade", "LAT": 25.833711, "LONG_": -80.166433, "DCODE": 1, "MORTALITY": "Human Related: Watercraft Collision", "created_user": "FWC", "created_date": "2021/10/15 11:30:55+00", "last_edited_user": "FWC", "last_edited_date": "2021/10/15 11:30:55+00"}]
Let's specify some columns, limit to 10 and add -t to output the results as a table.
sqlite-utils manatees.db -t \
"select REPDATE, MORTALITY, COUNTY from locations limit 10"
This outputs:
REPDATE MORTALITY COUNTY
---------------------- ----------------------------------- ------------
1974/04/03 00:00:00+00 Undetermined: Other Broward
1974/06/27 00:00:00+00 Human Related: Watercraft Collision Miami-Dade
1974/08/20 00:00:00+00 Human Related: Watercraft Collision Miami-Dade
1974/10/23 00:00:00+00 Human Related: Other Volusia
1974/11/11 00:00:00+00 Human Related: Other Brevard
1974/12/25 00:00:00+00 Human Related: Watercraft Collision Citrus
1974/12/27 00:00:00+00 Undetermined: Too Decomposed Indian River
1975/01/01 00:00:00+00 Verified: Not Necropsied Brevard
1975/01/19 00:00:00+00 Human Related: Watercraft Collision Miami-Dade
1975/01/31 00:00:00+00 Verified: Not Necropsied Brevard
Opening it in Datasette
Datasette provides a browser-based interface for exploring a SQLite database. Install that, (brew install datasette works for Homebrew users) and run it against your database:
datasette manatees.db
Then navigate to http://localhost:8001/ to start exploring your data.
Transforming the columns
Exploring the data helps identify improvements we can make to the way it is structured.
The LAT and LONG_ column look like they might be latitudes and longitudes. Renaming those columns to latitude and longitude will make that more obvious, and will also let us visualize the data using a Datasette plugin in a moment.
The created_user, last_edited_user and STATE columns aren't interesting: they always store the same value.
The X and Y columns are duplicates of the latitude and longitude.
Finally, the FIELDID column looks to be a unique identifier for every row in the database. This would make a good primary key for our table.
The sqlite-utils transform command can be used to apply transformations to a table - rename columns, dropping columns, assigning primary keys and more.
The following command will make a bunch of changes to the table structure in one go:
sqlite-utils transform manatees.db locations \
--rename LAT latitude \
--rename LONG_ longitude \
--drop created_user \
--drop last_edited_user \
--drop X \
--drop Y \
--drop STATE \
--drop OBJECTID \
--pk FIELDID
Now run sqlite-utils schema manatees.db to see the result of those changes:
CREATE TABLE "locations" (
[FIELDID] TEXT PRIMARY KEY,
[REPDATE] TEXT,
[REPYEAR] INTEGER,
[REPMONTH] INTEGER,
[REPDAY] INTEGER,
[SEX] TEXT,
[TLENGTH] FLOAT,
[COUNTY] TEXT,
[latitude] FLOAT,
[longitude] FLOAT,
[DCODE] INTEGER,
[MORTALITY] TEXT,
[created_date] TEXT,
[last_edited_date] TEXT
);
Visualizing the locations on a map
Datasette supports plugins, which enable extra features such as data visualizations.
The datasette-cluster-map plugin runs against any table with a latitude and longitude column, displaying them as points on a map. This is why we renamed those LAT and LONG_ columns earlier!
To install the plugin, quit the Datasette server (using Ctrl+C in the terminal window) and run the following:
datasette install datasette-cluster-map
Then start Datasette running again:
datasette manatees.db
If you visit http://localhost:8001/manatees/locations you will now see the first 1,000 rows as points on the map:
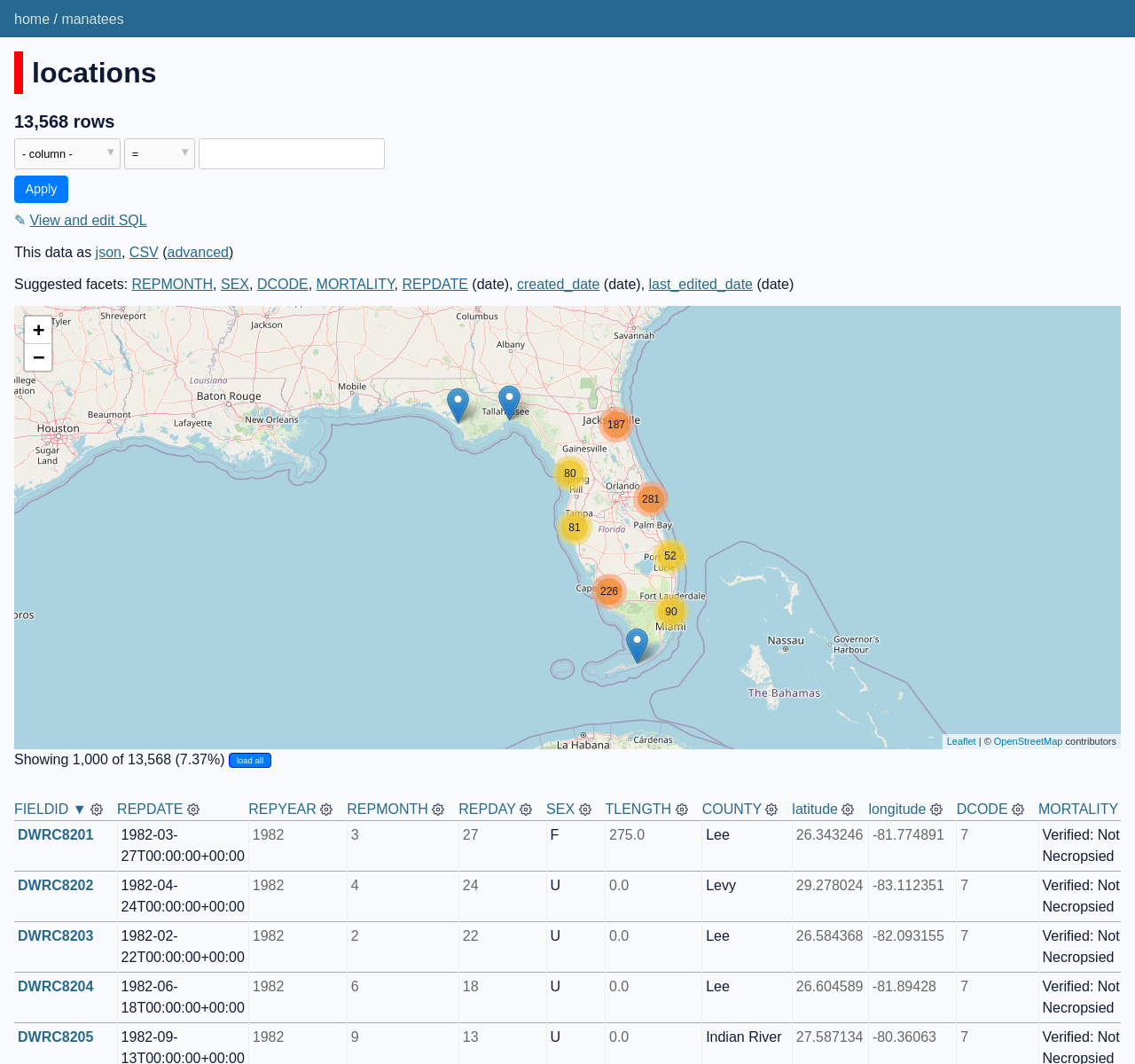
Click the "load all" button to load and display all 13,000+ locations.
Fixing the date columns
The date columns currently look like this:
2021/10/15 11:30:55+00
This is using YYYY/MM/DD format (at least it's not that confusing - for Europeans - American MM/DD/YYYY thing). I prefer the ISO standard format YYYY-MM-DD - so let's convert it.
The sqlite-utils transform command can be used to apply a transformation function to the values in one or more columns.
Here's how to use that to parse miscellaneous date and time formats and turn them into that standard ISO format:
sqlite-utils convert manatees.db locations \
REPDATE created_date last_edited_date \
'r.parsedatetime(value)'
This runs against the REPDATE, created_date and last_edited_date columns. It will use the built-in r.parsedatetime() recipe (documented here) to convert those values and save them back to the same column.
Run that command, then view the data in Datasette again. Those columns have now been converted!
Extracting columns into a separate table
There's one more improvement we will make to our data.
Open the table in Datasette, then use either the suggested facets list or the cog menu at the top of the columns to facet by both DCODE and MORTALITY.
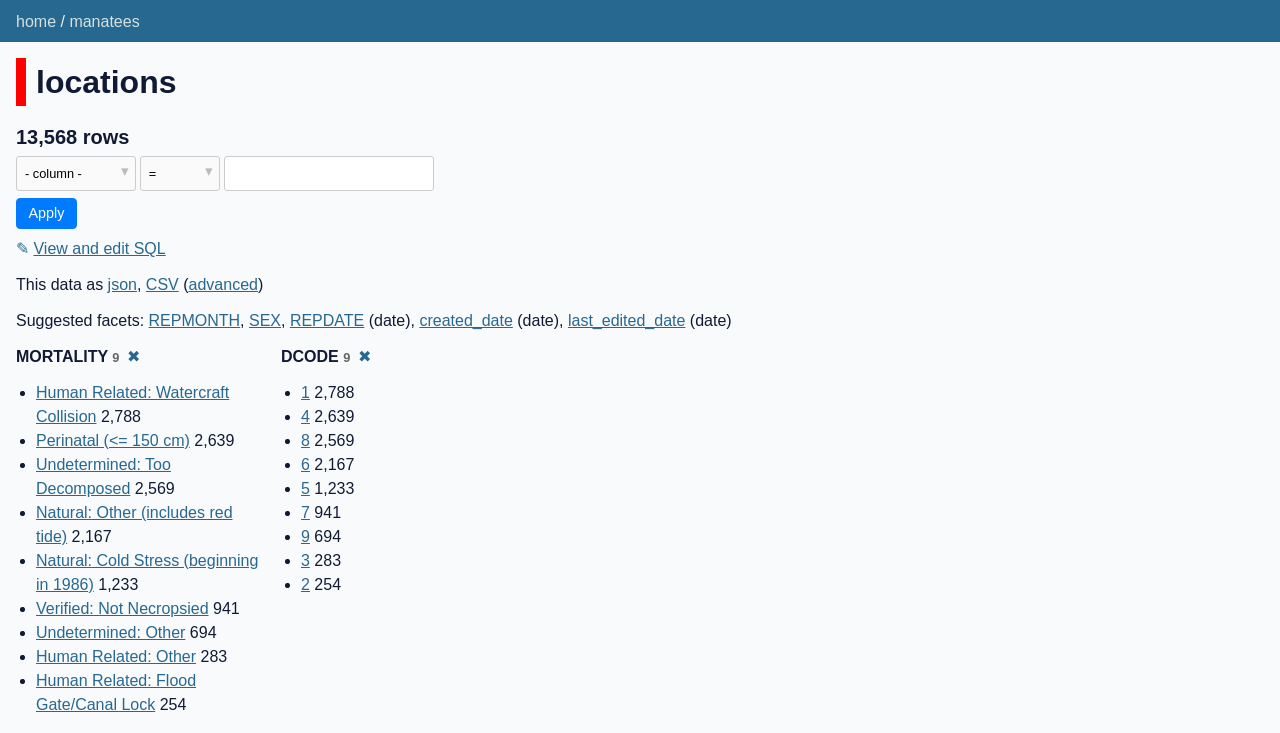
The result counts shown for the two facets are the same. This suggests that these two columns represent the same data - a DCODE of 1 corresponds to a MORTALITY of "Human Related: Watercraft Collision", 2 means "Human Related: Flood Gate/Canal Lock" and so on.
We can use the sqlite-utils extract command to extract these columns into a separate table.
sqlite-utils extract manatees.db locations DCODE MORTALITY \
--rename MORTALITY name \
--table mortality
Running this will create a new table called mortality and populate it with one row for each of the DCODE and MORTALITY pairs in the database. It removes those columns from the locations table and replaces them with a mortality_id foreign key column pointing to the new table.
Here's the output of sqlite-utils schema manatees.db showing the impact of those changes on the schema:
CREATE TABLE [mortality] (
[id] INTEGER PRIMARY KEY,
[DCODE] INTEGER,
[name] TEXT
);
CREATE TABLE "locations" (
[FIELDID] TEXT PRIMARY KEY,
[REPDATE] TEXT,
[REPYEAR] INTEGER,
[REPMONTH] INTEGER,
[REPDAY] INTEGER,
[SEX] TEXT,
[TLENGTH] FLOAT,
[COUNTY] TEXT,
[latitude] FLOAT,
[longitude] FLOAT,
[mortality_id] INTEGER,
[created_date] TEXT,
[last_edited_date] TEXT,
FOREIGN KEY([mortality_id]) REFERENCES [mortality]([id])
);
CREATE UNIQUE INDEX [idx_mortality_DCODE_name]
ON [mortality] ([DCODE], [name]);
Further reading
sqlite-utils offers a lot more tools for cleaning up data. Check out this documentation and these blog posts for more details.